Software
Plant DNA LLMs
Introduction
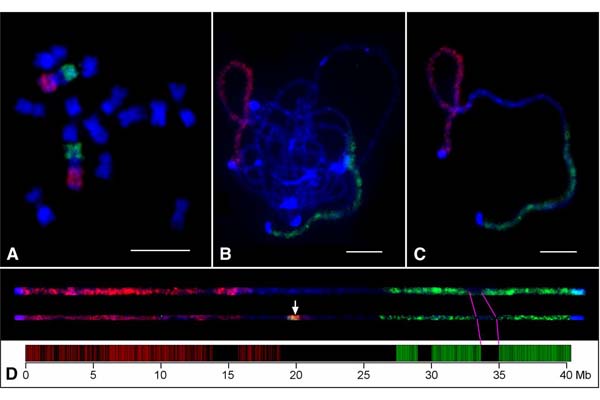
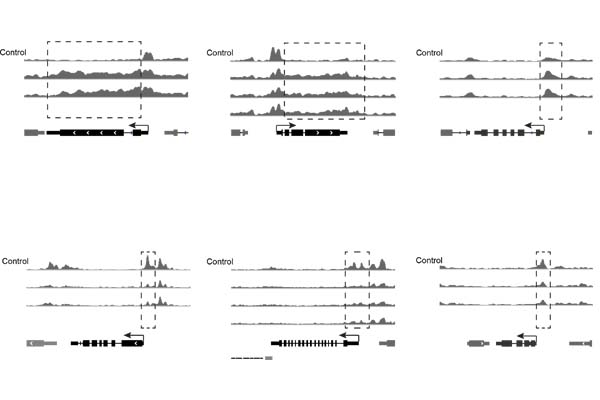
Chorus2
Design of genome-scale oligonucleotide-based probes for fluorescence in situ hybridization (FISH)
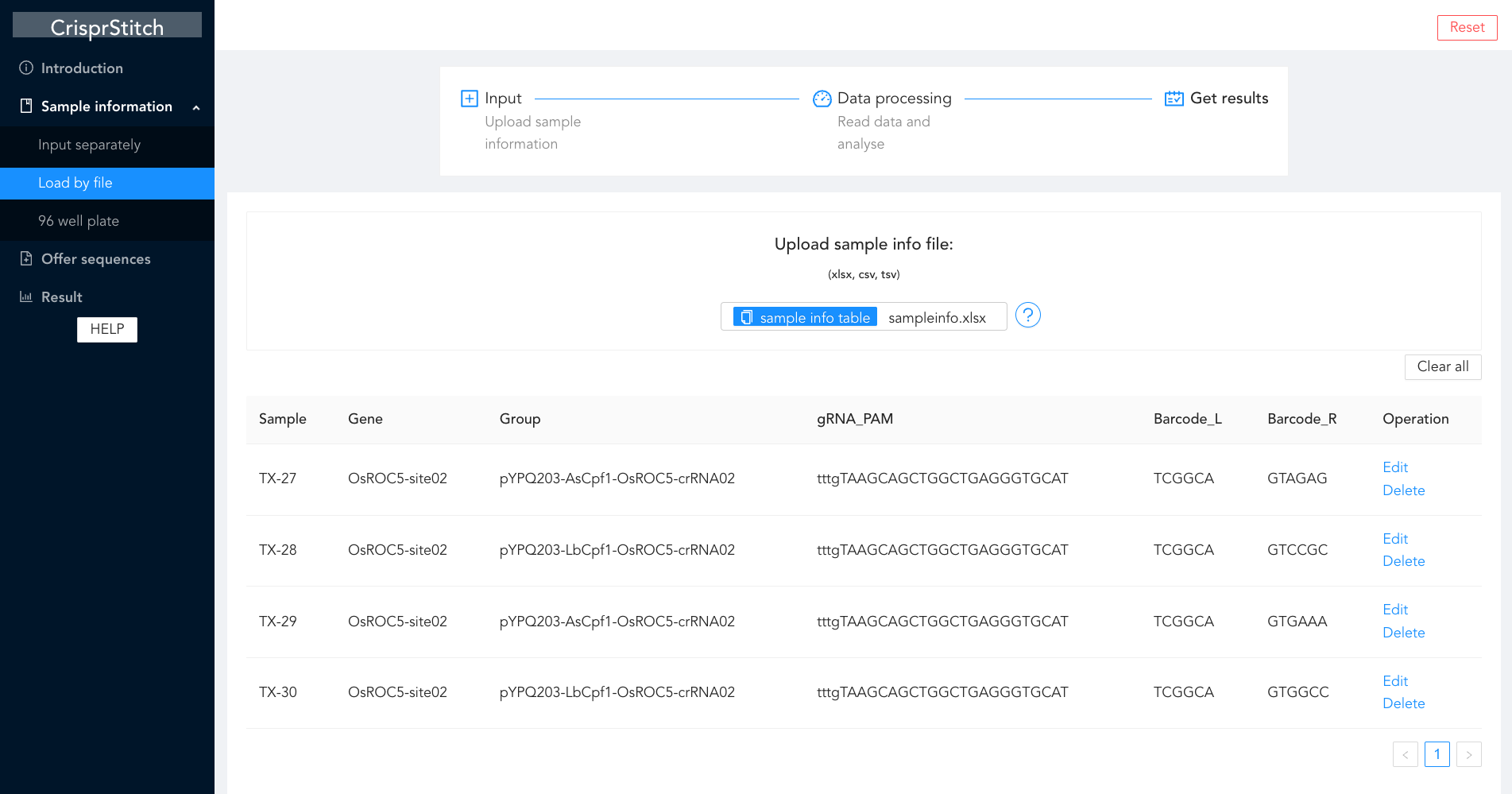
CrisprStitch
An integrated tool to fast evaluate the efficiency of CRISPR editing systems.
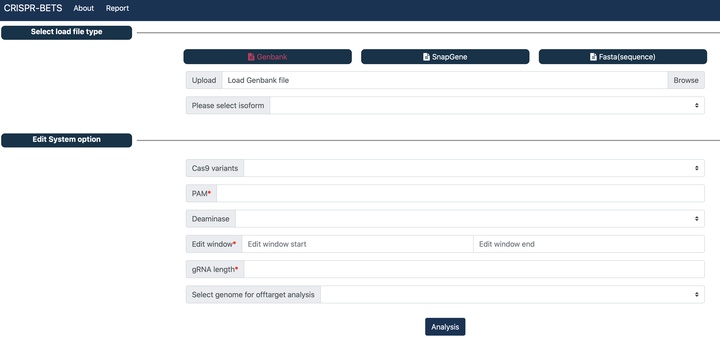
CRISPR-BETS
A gRNA design software for base editing knockout using stop codon.
Popera
A software specially designed for DHS identification. Github Introduction DNase I hypersensitive sites (DHSs) are regions of chromatin that are sensitive to cleavage by the...
Latest
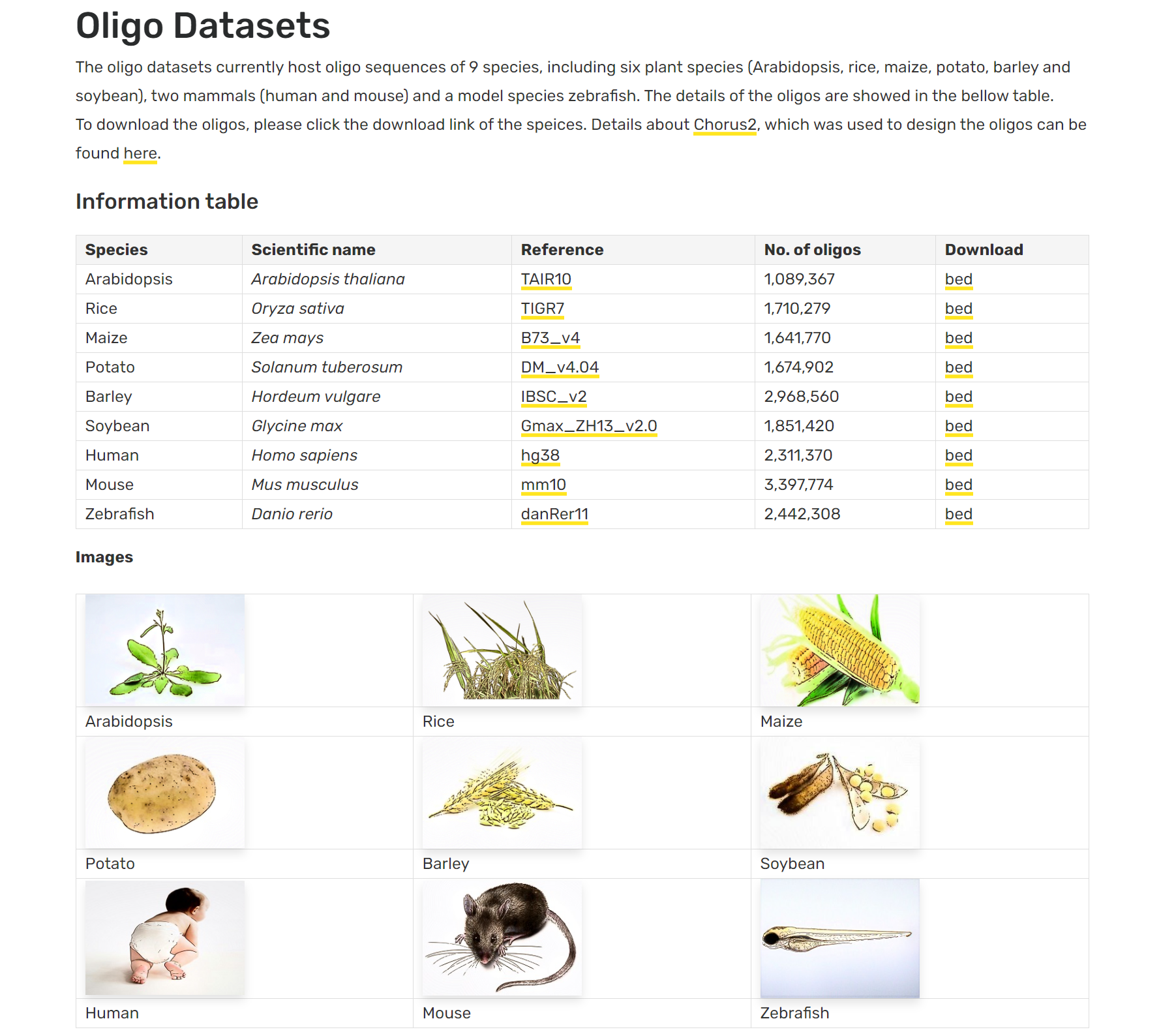
Chorus2 oligo datasets
Oligo datasets which host oligo sequences for Oligo-FISH. Details Introduction The oligo datasets currently host oligo sequences of nine species, including Arabidopsis, rice, maize, potato, barley, soybean, human, mouse and...
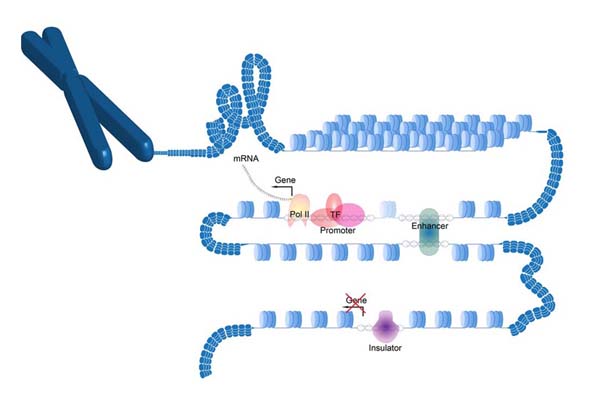
PlantDHS
PlantDHS: a database for DNase I hypersensitive sites (DHSs) in plants.
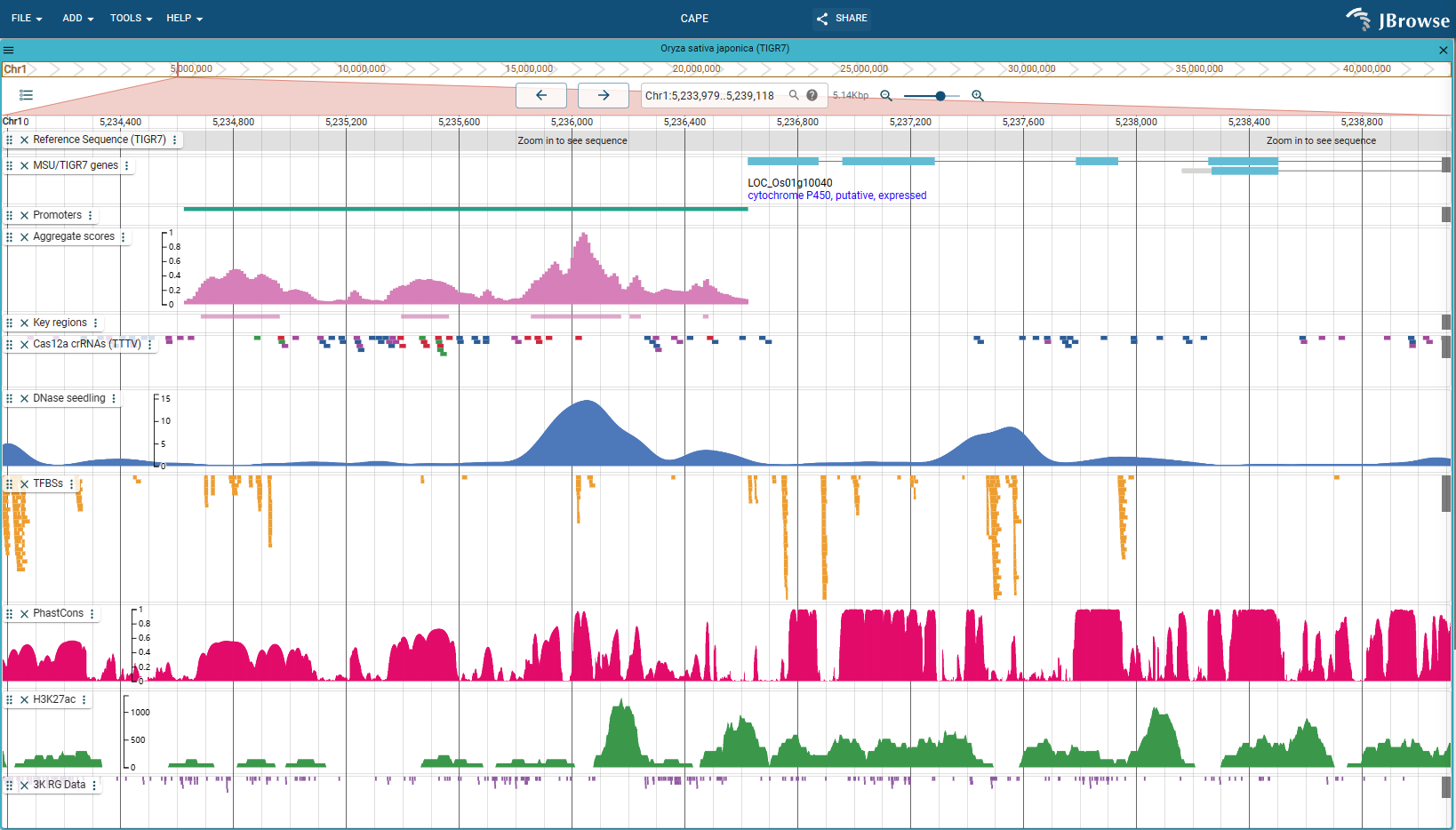
CAPE
CAPE: An efficient CRISPR-Cas12a promoter editing system for crop improvement [JBrowse preview](https://bioinfor.yzu.edu.cn/jbrowse2/cape) ## Introduction we demonstrate a **CAPE** based promoter editing and tuning pipeline for efficient production of useful quantitative...

CAPE
CAPE: An efficient CRISPR-Cas12a promoter editing system for crop improvement JBrowse preview Introduction we demonstrate a CAPE based promoter editing and tuning pipeline for efficient production of useful quantitative trait...
Lab meeting

Splicing-mediated activation of SHAGGY-like kinases underpinning carbon partitioning in Arabidopsis seeds
Abstract: Glycogen synthase kinase 3 (GSK3) family members serve as signaling hubs for plant development and stress responses, yet the underlying mechanism of their transcriptional regulation remains a long-standing mystery....

Role of lncRNAs in cis- and trans-regulatory responses to salt in Populus trichocarpa
Abstract: Long non-coding RNAs (lncRNAs) are emerging as versatile regulators in diverse biological processes. How- ever, little is known about their cis- and trans-regulatory contributions in gene expression under salt...

ReFeaFi: Genome-wide prediction of regulatory elements driving transcription initiation
Abstract: Regulatory elements control gene expression through transcription initiation (promoters) and by enhancing transcription at distant regions (enhancers). Accurate identification of reg- ulatory elements is fundamental for annotating genomes and...

Chromatin accessibility landscape and regulatory network of high-altitude hypoxia adaptation
Abstract: High-altitude adaptation of Tibetans represents a remarkable case of natural selection during recent human evolution. Previous genome-wide scans found many non-coding variants under selection, suggesting a pressing need to...

DNA methylation-free Arabidopsis reveals crucial roles of DNA methylation in regulating gene expression and development
Abstract: A contribution of DNA methylation to defense against invading nucleic acids and maintenance of genome integrity is uncontested; however, our understanding of the extent of involvement of this epigenetic...

ReFeaFi: Genome-wide prediction of regulatory elements driving transcription initiation
Abstract: Regulatory elements control gene expression through transcription initiation (promoters) and by enhancing transcription at distant regions (enhancers). Accurate identification of regulatory elements is fundamental for annotating genomes and understanding...

A comprehensive catalog of predicted functional upstream open reading frames in humans
Abstract: Upstream open reading frames (uORFs) latent in mRNA transcripts are thought to modify translation of coding sequences by altering ribosome activity. Not all uORFs are thought to be active...

ALK, the Key Gene for Gelatinization Temperature, is a Modifier Gene for Gel Consistency in Rice
Abstract: Gelatinization temperature (GT) is an important parameter in evaluating the cooking and eating quality of rice. Indeed, the phenotype, biochemistry and inheritance of GT have been widely studied in...

Linker histones are fine-scale chromatin architects modulating developmental decisions in Arabidopsis
Abstract: Background: Chromatin provides a tunable platform for gene expression control. Besides the well-studied core nucleosome, H1 linker histones are abundant chromatin components with intrinsic potential to influence chromatin function....